Reproducible scientific manuscripts
R Markdown and manuscripts
Of the many great resources for creating reproducible documents in R Markdown, two packages stand out: bookdown and papaja.
bookdown is a powerful tool for creating not only books, but also other documents that cross-reference tables, figures, and sections/chapters.
papaja is specifically designed for APA-formatted documents and includes functions for creating tables and figures.
Although many researchers rely on Word-based work flows, I found the existing resources for combining R Markdown and Microsoft Word to be limited.
The documentation for bookdown is clear and thorough, but privileges the more flexible and easily customized HTML and PDF outputs over Word outputs.
papaja also privileges PDF over Word outputs in its documentation and functionality.
As a result, these packages fall short of providing the flexible and customizable Word outputs needed for writing scientific manuscripts.
The work-around approach—creating PDF outputs and converting the underlying .tex files to Word with Pandoc—also produces less than satisfactory results.
I wanted to create Microsoft Word documents using R Markdown that (a) already had the formatting I needed without a lot of post-output effort and (b) were also easily customized depending on the journal/audience.
Thus, I combined the strengths of bookdown and papaja into the flexible template described below.
The template
I developed this template for creating scientific manuscripts to take advantage of R Markdown’s reproducibility, Word’s ease of collaboration, and bookdown’s cross-referencing capabilities.
Organization
The R Markdown files and R scripts for the template are organized in a hierarchical fashion. This structure keeps the information tidy and concise at each stage, from data wrangling, analysis, and visualization to presentation/writing.
There are separate R Markdown files for each section:
- Title page
- Abstract
- Introduction
- Methods
- Results
- Conclusion/discussion
- Acknowledgments/funding
There is more information about using child documents and source code in my R Markdown Guide.
Formatting
The formatting can be easily customized by changing the Word styles in the template.docx file or editing the fenced code blocks in the R Markdown files.
The template file tells Pandoc how to convert from markdown text to Word output. Pandoc automatically assigns particular types of text to particular Word styles; for example, first level headers in R Markdown receive the Header 1 format in Word, as shown in the image below. These Word styles are defined in the template document.
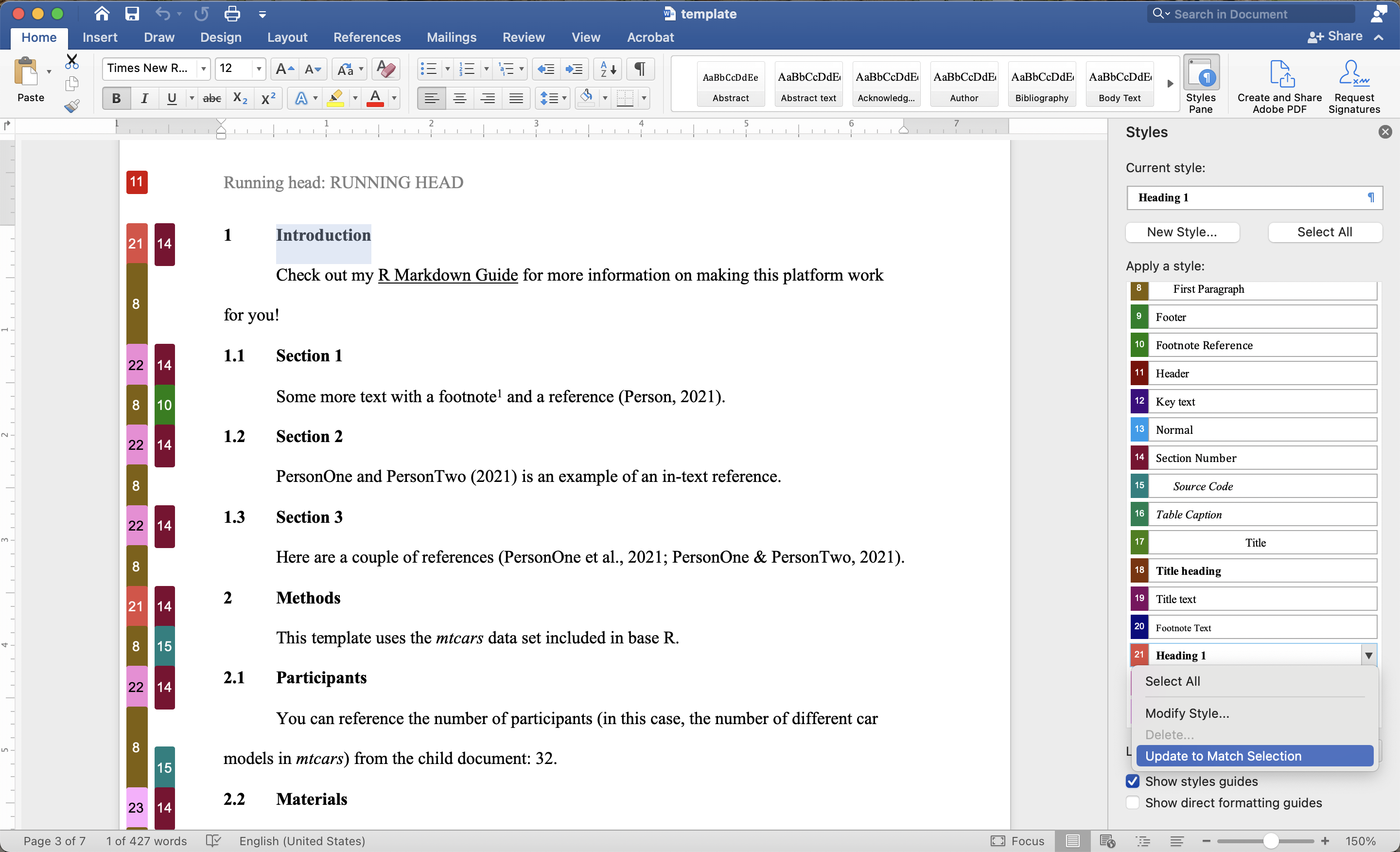
In order to edit these styles:
- Open the template document
- Open the Styles Pane
- Highlight the text whose format you want to change
- Make the desired changes (e.g., italicizing Header 1)
- Locate the format in the Styles Pane
- Select Update to Match Selection from the drop-down menu
If you want to specify exactly which Word style a chunk of text should receive, you can use fenced code blocks. Below, I specify that I want the Abstract text format, which I defined in my template file, to apply to the text between the three colons.
---
output: bookdown::word_document2
---
# Abstract {-}
::: {custom-style="Abstract text"}
Your abstract goes here.
:::
Limitations
There are two main limitations to using Word outputs. One is that, to my knowledge, there is no way to include line numbers or update headers automatically. This will need to be done in the Word output.
More importantly, there is currently no way easy way to incorporate track-changes from collaborators. I recommend using Git to compare your document to your collaborators’ or using the Combine Documents functionality in Word. Unfortunately, neither of these solutions updates your actual R Markdown documents. This will need to be done by hand if needed.
]]>